mclUMI
mclUMI is an open-source, modular, and scalable Python programming interface that
possesses multiple modules for improving sequencing accuracy.
mclUMI offers read I/O, preprocessing, UMI deduplication, and
chimeric read removal based on homotrimer blocks.
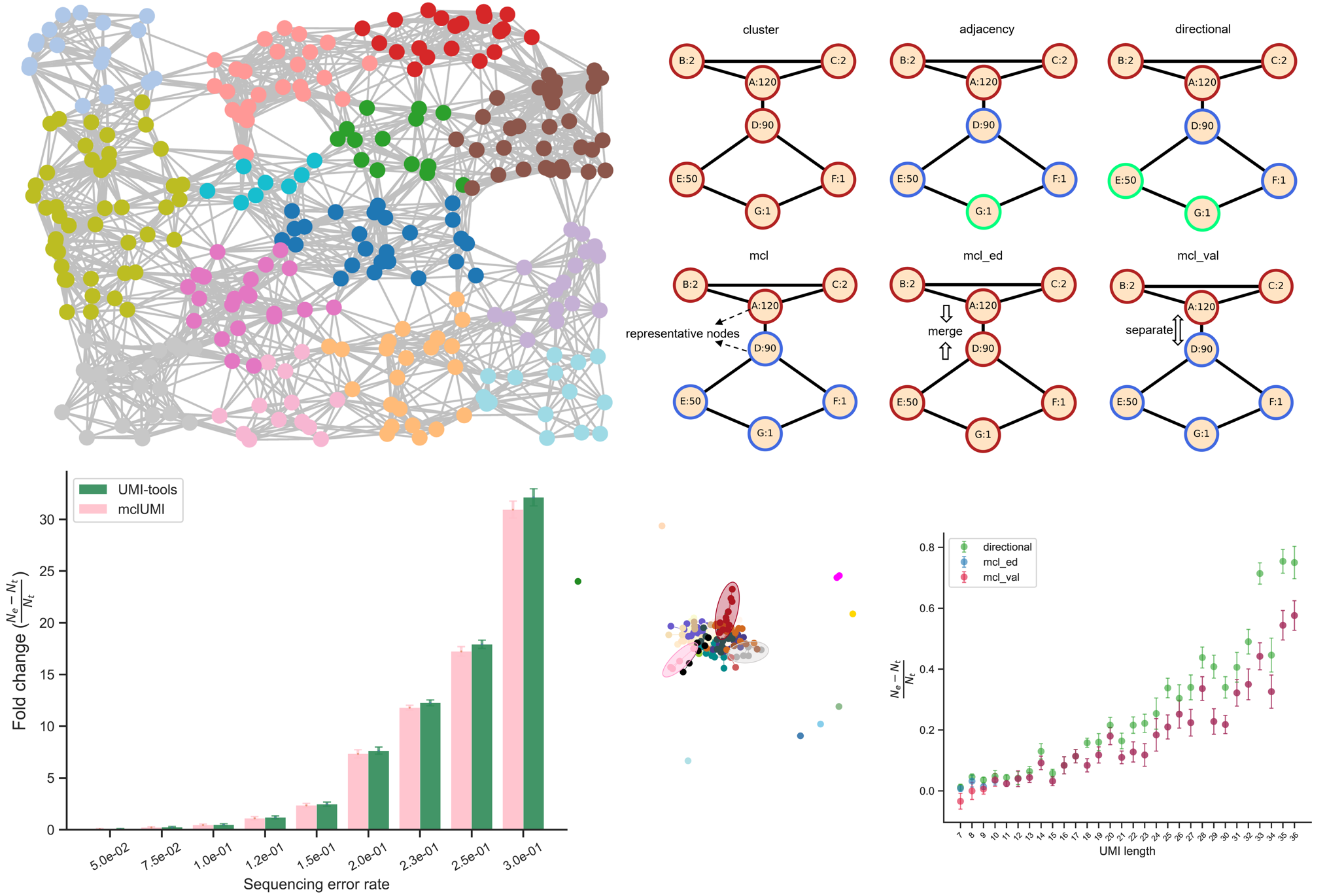
Intriguingly, it utilizes the Markov clustering algorithm to allow us to gain
multiple choices of UMI count matrices, which is different from currently
available algorithms or programs. This avoids one-size-fits-all strategies for generating
deduplicated UMI counts and can help us find the best solution through its built-in expansion and inflation
settings for reads that sequenced by extremely error-prone or accurate sequencing technologies.
mclUMI strives to make read quantification more accurate and easier
and will accelerate the biological translation.